The edman degradation process is used to determine the sequence of a protein. So after you purified your protein, confirmed that you are working with one sequence, and reduce the protein sequence into smaller fragments (if protein is very large you must break it into smaller fragments), then you are ready to sequence your protein?
What is protein sequencing?
Since proteins are composed of amino acids (20 amino acids) it is imperative to determine the sequence to understand the function of the protein.
So, how does the Edman degradation process work?
The process is quite simple actually.
Take your protein fragment and run it through repeated cycles with a reagent called phenylisothiocyanate (also known as PITC).
What this reagent does is it reacts with the N-terminal of an amino group under mildly alkaline conditions to form a phenylthiocarbamyl (PTC) adduct.
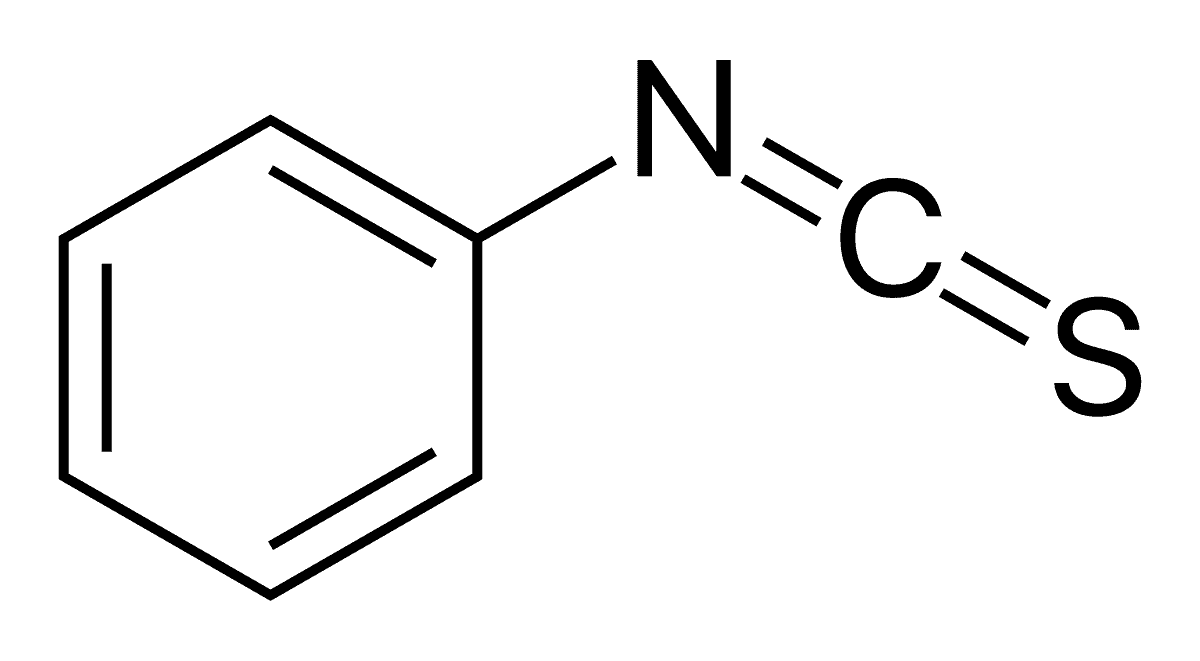
Structure of PITC
This product is then treated with anhydrou trifluoroacetic acid which breaks away the N-terminal residue from the other amino acids (without messing up the residues of the other amino acids in the polypeptide fragment) and then can be detected after treated with aqueous acid (so that the structure becomes more stable)
The mechanism is shown below:
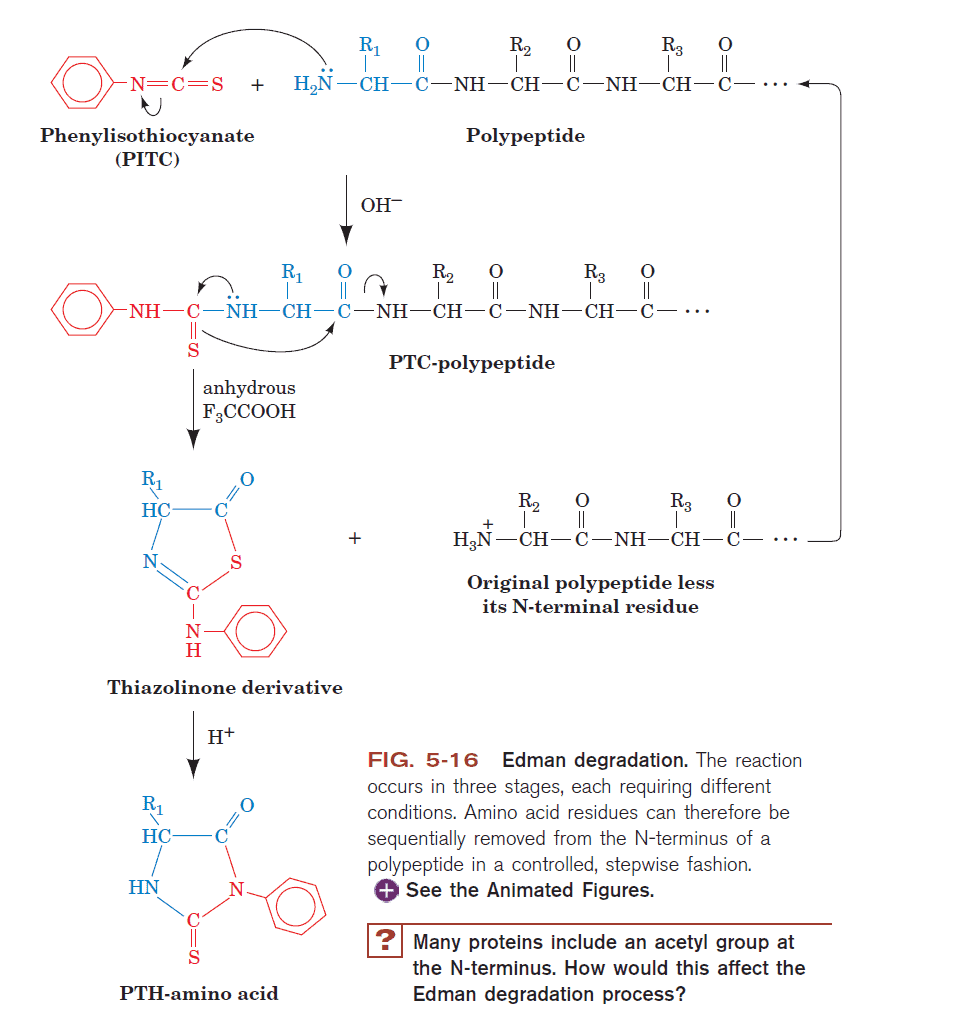
Source: Voet,Voet, and Pratt Biochemistry Textbook
If you have any questions or would like to share your reviews on the Edman Degradation Process, then comment down below. I would love to hear what you have to think.
